OXOID培养基 SR0082E 鲍特菌选择性添加剂(头孢力新) BORDETELLA SELECTIVE SUPP. 454.5497142144
日度归档:2026年4月15日
Whatman 滤膜 10549663 BA79 64MMx400MM PK/10 LAM
Whatman 滤膜 10549663 BA79 64MMx400MM PK/10 LAM BA79 64MMx400MM PK/10 LAM
nunc 130193 BioLite 75cm2 Flask, Plug Seal
nunc 130193 BioLite 75cm2 Flask, Plug Seal
NISSUI 日本日水培养基 06054 Food Stamp “Nissui” TCBS Agar (TCBS)
Whatman 滤膜 2812-259 二氧化硅套管, 锥形, 25 x 90mm/10
Whatman 滤膜 2812-259 二氧化硅套管, 锥形, 25 x 90mm/10 THIMBLE SILICA 25x90MM 10/PK
白藜芦醇及其相关产品
| 产品编号 | 产品名称 | 产品规格 | 产品等级 | 产品价格 |
| 184-02771 | Resveratrol, Synthetic 白藜芦醇 |
1G | Wako Special Grade | – |
| 180-02773 | Resveratrol, Synthetic 白藜芦醇 |
5G | Wako Special Grade | – |
| 182-02772 | Resveratrol, Synthetic 白藜芦醇 |
25G | Wako Special Grade | – |
| 070-06141 | Gnetin C Standard 反式白藜芦醇二聚体标准品 |
50mg | for Food Analysis | – |
| 226-02021 | ε-Viniferin Standard ε-葡萄素标准品 |
20mg | for Food Analysis | – |
白藜芦醇及其相关产品
白藜芦醇,一种在葡萄皮和其他可食用的植物和相关的食物产品中发现的多酚,是一种抗氧化剂。最近,各种白藜芦醇有益于健康的好处已经被广泛研究。我们还供应反式白藜芦醇二聚体和ε‐葡萄素标准品,即白藜芦醇二聚体。
◆白藜芦醇
|
中文名称 |
英文名称 |
等级 |
包装(g) |
货号 |
储存条件 |
|
白藜芦醇 |
Resveratrol,Synthetic |
和光特级 |
1 |
184-02771 |
‐20 ˚C保存 |
|
5 |
180-02773 |
||||
|
25 |
18202772 |
检测:98±%(HPLC)
外观:白色~浅棕色,结晶性粉末~粉末状或块状
化学名称:5‐[(1)2(4‐‐‐羟基苯基)乙烯基]‐‐1,3-苯二酚
CAS NO.:501-36-0
化学结构式:
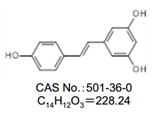
◆反式白藜芦醇二聚体标准品
|
中文名称 |
英文名称 |
等级 |
包装(mg) |
货号 |
储存条件 |
|
反式白藜芦醇二聚体 |
Gnetin C Standard |
食品分析用 |
50mg |
070-06141 |
‐20 ˚C保存 |
检测:97±%(HPLC)
外观:白色~土黄色,结晶性粉末~粉末或者块状
化学名称:REL‐5‐[(2 R,3 R)‐2,3‐二氢‐4‐羟基‐2‐(4‐羟基苯基)‐6‐[(1)2(4‐‐‐羟基苯基)乙烯基]苯并呋喃基]‐3‐‐1,3‐苯二酚
注:反式白藜芦醇暴露于荧光或自然光时,可能会分解
CAS NO.:84870-54-2
化学结构式:

◆ε‐葡萄素标准品
|
中文名称 |
英文名称 |
等级 |
包装(mg) |
货号 |
储存条件 |
|
ε‐葡萄素标准品 |
ε‐Viniferin Standard |
食品分析用 |
20mg |
226-02021 |
‐20 ˚C保存 |
检测:98±%(HPLC)
外观:白色~浅灰棕色,结晶性粉末~粉末或者块状
化学名称:5‐[(2R,3R)‐2,3‐二氢‐6‐羟基‐2‐(4‐羟基苯基)‐4‐[(1E)‐2‐(4‐羟基苯基)乙烯基]苯并呋喃基]‐3‐‐1,3‐苯二酚
注:反式白藜芦醇二聚体包含在绿芽和葡萄藤中
CAS NO.:62218-08-0
化学结构式:
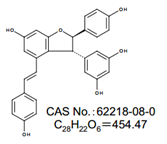
◆产品列表
|
产品编号 |
产品名称 |
产品规格 |
|
184-02771 |
Resveratrol, Synthetic 白藜芦醇 |
1G |
|
180-02773 |
Resveratrol, Synthetic 白藜芦醇 |
5G |
|
182-02772 |
Resveratrol, Synthetic 白藜芦醇 |
25G |
|
070-06141 |
Gnetin C Standar d反式白藜芦醇二聚体标准品 |
50mg |
|
226-02021 |
ε-Viniferin Standar dε-葡萄素标准品 |
20mg |
◆相关产品
|
产品编号 |
产品名称 |
产品规格 |
|
182-02831 |
Resveratrol Standard 白藜芦醇标准品 |
100 mg |
◆其他食品分析用标准品
|
中文名称 |
英文名称 |
等级 |
包装 |
货号 |
储存条件 |
备注 |
|
薯蓣皂素标准品 |
Diosgenin Std., 98.0+%(HPLC) |
食品分析用 |
100mg |
043-32621 |
2~10℃保存 |
一种含在山药中的孕酮同源物 |
|
白藜芦醇 |
Resvertrol, 98.0+%(HPLC) |
生物化学用 |
100mg |
185-01721 |
‐20 ˚C保存 |
自然提取 |
|
500mg |
181-01723 |
|||||
|
杜克甙A标准品 |
Dulcoside Std, 98.0+%(HPLC,) |
食品分析用 |
25mg |
048-31211 |
2~10℃保存 |
甜叶菊中甜味剂
|
|
100mg |
044-31213 |
|||||
|
异甜菊苷标准品 |
Isosteviol Std., 98.0+%(HPLC,) |
测定甜菊糖用 |
1g |
098-05681 |
2~10℃保存 |
|
|
莱苞迪甙A标准品 |
Rebaudioside A Std., |
食品分析用 |
100mg |
189-02581 |
2~10℃保存 |
|
|
1g |
185-02583 |
|||||
|
莱苞迪甙B标准品 |
Rebaudioside B Std., |
食品分析用 |
25mg |
188-02551 |
2~10℃保存 |
|
|
100mg |
184-02553 |
|||||
|
莱苞迪苷C |
Rebaudioside C, 90.0+% (HPLC) |
食品分析用 |
25mg |
181-02541 |
2~10℃保存 |
|
|
莱苞迪苷D |
Rebaudioside D |
食品分析用 |
5mg |
180-02511 |
‐20 ˚C保存 |
|
|
莱苞迪苷F |
Rebaudioside F |
食品分析用 |
5mg |
186-02611 |
2~10℃保存 |
|
|
甜茶苷标准品 |
Rubusoside Std., |
食品分析用 |
25mg |
187-02521 |
‐20 ˚C保存 |
|
|
100mg |
183-02523 |
|||||
|
甜菊糖标准品 |
Steviol Std., |
食品分析用 |
25mg |
192-15701 |
2~10℃保存 |
|
|
100mg |
198-15703 |
|||||
|
甜菊双糖甙标准品 |
Steviolbioside Std., |
食品分析用 |
25mg |
199-15691 |
2~10℃保存 |
|
|
100mg |
195-15693 |
|||||
|
甜叶菊甙元标准品 |
Stevioside Std., 99.0+ % (HPLC) |
食品分析用 |
100mg |
199-16291 |
2~10℃保存 |
100bp Ladder DNA marker 美国AXYGEN
名称:100bp Ladder DNA marker
品牌:美国AXYGEN
订货号:
![]()
咨询此产品
产品介绍
片段长度:100bp-2000bp
|
产品序号 |
规格 |
|
|
M-DNA-100bp |
50ug/500ul |
|
Whatman 滤膜 29180120 BENCHKOTE REEL 92CMx50M 1/PK, PROMO
Whatman 滤膜 29180120 BENCHKOTE REEL 92CMx50M 1/PK, PROMO BENCHKOTE REEL 92CMx50M 1/PK, PROMO
NISSUI 日本日水培养基 05009 亚硒酸盐胱胺酸基础培养基 Selenite Cystine Broth Base 选择增加沙门氏菌 100g
Whatman 滤膜 1450-6955 GR 50 750MMx1000M REELS
Whatman 滤膜 1450-6955 GR 50 750MMx1000M REELS GR 50 750MMx1000M REELS
Pall颇尔滤膜 4272 FILTER AQUAPREP-V 50MM PK-20
Pall颇尔滤膜 4272 FILTER AQUAPREP-V 50MM PK-20 646 5172
Whatman 滤膜 112804 NUC PC 293MM 0.08uM 25/PK
Whatman 滤膜 112804 NUC PC 293MM 0.08uM 25/PK NUC PC 293MM 0.08uM 25/PK
Pall颇尔滤膜 OD010C35 DEVICE CENTRIFU NANO 10K PK500
Pall颇尔滤膜 OD010C35 DEVICE CENTRIFU NANO 10K PK500 1343 10745
地衣淀粉/地衣多糖 [冰岛苔] Lichenan (Icelandic Moss) 4g 货号:P-LICHN Megazyme中文站
地衣淀粉/地衣多糖 [冰岛苔]
英文名:Lichenan (Icelandic Moss) 4g
货号:P-LICHN
规格:4 grams
暂无详情描述
暂无问题解答
暂无视频
Whatman 滤膜 10300140 GR 589/2 41.5MM 100/PK
Whatman 滤膜 10300140 GR 589/2 41.5MM 100/PK GR 589/2 41.5MM 100/PK
Lumiprobe品牌代理商
Lumiprobe品牌代理商–上海金畔
1. Lumiprobe品牌介绍
Lumiprobe Corporation是美国一家高品质生物技术公司,专业提供分子生物学研究用的活
性荧光染料。从2006年开始,公司生产并销售生命科学研究和诊断学应用的优质化学药品。
产品主要有:活性染料(Reactive Dye)和SYBR Green I 染料,用于寡核苷酸合成的亚
磷酰胺,点击化学和其它试剂。产品主要应用:点击化学(Click Chemistry)、蛋白质组学
研究中的双向荧光差异凝胶电泳(2D DIGE)和实时荧光定量PCR(Realtime PCR)。产品包括
:N-羟基琥珀酰亚胺酯类活性染料;羰基活性荧光染料;氨基类染料 ;巯基反应性染料;
羧酸类染料。
2. Lumiprobe重点产品
N-羟基琥珀酰亚胺酯类活性染料;羰基活性荧光染料;氨基类染料 ;巯基反应性染料;羧
酸类染料。
3. Lumiprobe官网
http://www.lumiprobe.com/
4. 金畔生物代理Lumiprobe品牌联系方式
上海金畔生物科技有限公司
地 址: 上海市浦东新区东靖路699弄36栋701室
邮 编: 201208
qq: 2743691513
固话总机:021-50837765
订货热线:15221999938
网 址: www.jinpanbio.com
Email:sales@jinpanbio.com
Pall颇尔滤膜 OA030C12 CAPSULE MINIMATE TFF 30K PK1
Pall颇尔滤膜 OA030C12 CAPSULE MINIMATE TFF 30K PK1 400 3197
Wako 和光纯药 SuperSep™ Phos-tag™ (50μmol/l), 6%, 13well Phos-tag™ 预制胶50μmol/l,6%,13孔
Wako 和光纯药 SuperSep™ Phos-tag™ (50μmol/l), 6%, 13well Phos-tag™ 预制胶50μmol/l,6%,13孔
Wako Wako ——————和光纯药 192-17401
SuperSep™ Phos-tag™ (50μmol/l), 6%, 13well
Phos-tag™ 预制胶50μmol/l,6%,13孔
– 5块 3,370.00 -Wako ——————和光纯药 193-16711
SuperSep™ Phos-tag™ (50 μmol/L), 10%,13 well
Phos-tag™ 预制胶50μmol/l,10%,13孔
for Electrophoresis 5 EA 3,370.00 (9003-05-8)Wako ——————和光纯药 195-17371
SuperSep™ Phos-tag™ (50μmol/l), 7.5%, 13well
Phos-tag™ 预制胶50μmol/l,7.5%,13孔
– 5块 3,370.00 -Wako ——————和光纯药 199-17391
SuperSep™ Phos-tag™ (50μmol/l), 6%, 17well
Phos-tag™ 预制胶50μmol/l,6%,17孔
– 5块 3,370.00 -Wako ——————和光纯药 196-16701
SuperSep™ Phos-tag™ (50 μmol/L), 15%,17 well
Phos-tag™ 预制胶50μmol/l,15%,17孔
– 5块 3,370.00 -Nard ——————- 300-93523
Phos-tag™ Acrylamide AAL-107
Phos-tag™ 丙烯酰胺
– 2 mg 3,060.00
重组无细胞蛋白合成系统
| 产品编号 | 产品名称 | 产品规格 | 产品等级 | 产品价格 |
| GFK-PF201-0.25-EX | PUREfrex® 2.0 | 1KIT | - | - |
| GFK-PF201-0.25-5-EX | PUREfrex® 2.0 | 1KIT | - | - |
| GFK-PF213-0.25-EX | PUREfrex® 2.1 | 1KIT | - | - |
| GFK-PF213-0.25-5-EX | PUREfrex® 2.1 | 1KIT | - | - |
| GFK-PF003-0.5-EX | DnaK Mix | 1KIT | - | - |
| GFK-PF004-0.5-EX | GroE Mix | 1KIT | ||
| GFK-PF005-0.5-EX | DS supplement | 1KIT |
重组合无细胞蛋白合成系统![]()
PUREfrex® 2.0
◆简介
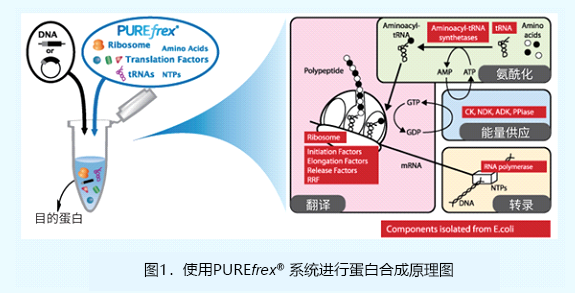
PUREfrex® 试剂盒是在东京大学的Takuya Ueda教授所发明的PUREsystem技术基础上,新开发的一款重组合无细胞蛋白合成试剂盒。
反应系统由蛋白质、核糖体、氨基酸和NTPs组成[1,2],其中蛋白行使转录、翻译和能量供应的功能。蛋白与核糖体为分别单独高度纯化后,再重新组合成蛋白合成系统,而非直接从大肠杆菌S30中提取。当合成蛋白时,仅需将编码目的蛋白的模板DNA或mRNA添加到反应混合液中并孵育数小时,即可完成反应。本系统最大的特色,是在体外以转录相关因子重新组合一套表达系统,并可根据需要来调整反应混合物的成分,而不必担心高背景会影响下游的应用。进行蛋白表达仅需将编码目标蛋白的模板DNA或mRNA加入到反应体系中,然后孵育2-4小时即可完成反应。PUREfrex® 试剂盒的所有蛋白组分均不带标签,因此目的蛋白可融合任意标签进行纯化和检测。
>>>无细胞表达的优势<<<
● 无需制备克隆
● 无需考虑培养条件
● 无需考虑表达所需的诱导条件
● 来源于宿主的污染少
◆PUREfrex® 系列
● PUREfrex® 1.0 第一代产品
● PUREfrex® 2.0 第二代产品,表达量更高,污染水平更低;
● RNA酶与β-半乳糖苷酶污染大大降低;
● 每1 µL反应混合物中的脂多糖(LPS)低于0.1 EU。
● PUREfrex® 2.1 更适合二硫键的形成
☆升级至PUREfrex® 2.0
1.合成原核和真核蛋白的结果显示,用PUREfrex® 2.0合成时,各种蛋白的合成量增加。
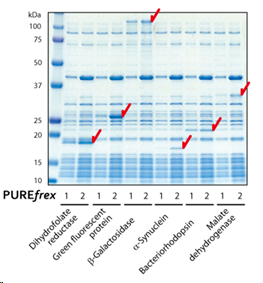
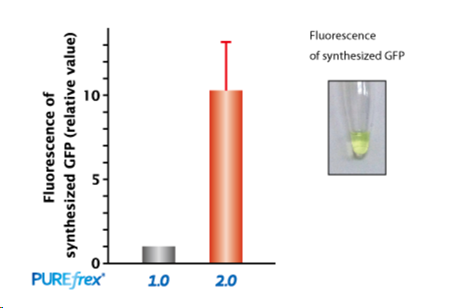
2.GFP蛋白合成的结果显示,用PUREfrex® 2.0合成时,可以观察到荧光强度增强了
2.10倍以上(每单位反应产物)。
3.合成需要形成二硫键(SS键)的大肠杆菌酸性磷酸酶(AppA1)时,在PUREfrex® 2.0基础上,
3.添加了DS supplement的结果显示,存在氧化剂和二硫键异构酶时,活性蛋白合成量增加。
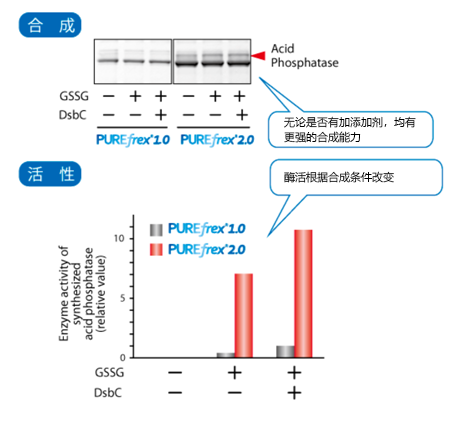
1:AppA有5个二硫键,是其中一个位点在不连续的半胱氨酸之间存在的二硫键。
4.正确高级结构蛋白的合成结果显示,存在分子伴侣2的情况下用PUREfrex® 2.0,蛋白合成量增加。
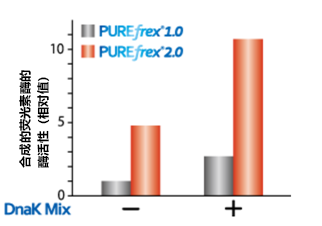
2 DnaK Mix:DnaK / DnaJ / GrpE mixture为配套分子伴侣
◆特点
● 可以同时加入多种模板进行反应,以合成Fab(带二硫键)及多聚体等带二级结构的多肽
● 可合成活细胞难以合成的强毒性蛋白
● 可直接使用PCR产物来作为模板DNA
● 单位体积内合成的蛋白量几乎恒定,不随反应体积变化而产生显著差异
● 操作简便,仅需在37℃孵育数小时
● 可以合成带标签的蛋白用于下游纯化和检测
● 产品经优化升级,合成量大大提高
◆应用
制备目的蛋白
● 原核蛋白
● 真核蛋白
● 膜蛋白
● 二硫键蛋白
● 含有非天然氨基酸的蛋白质等
蛋白基础研究
● 翻译
● 蛋白合成后折叠
体外展示技术
● 核糖体展示技术
● mRNA展示技术
◆应用实例
利用PUREfrex® 系统合成并一步纯化DHFR-His
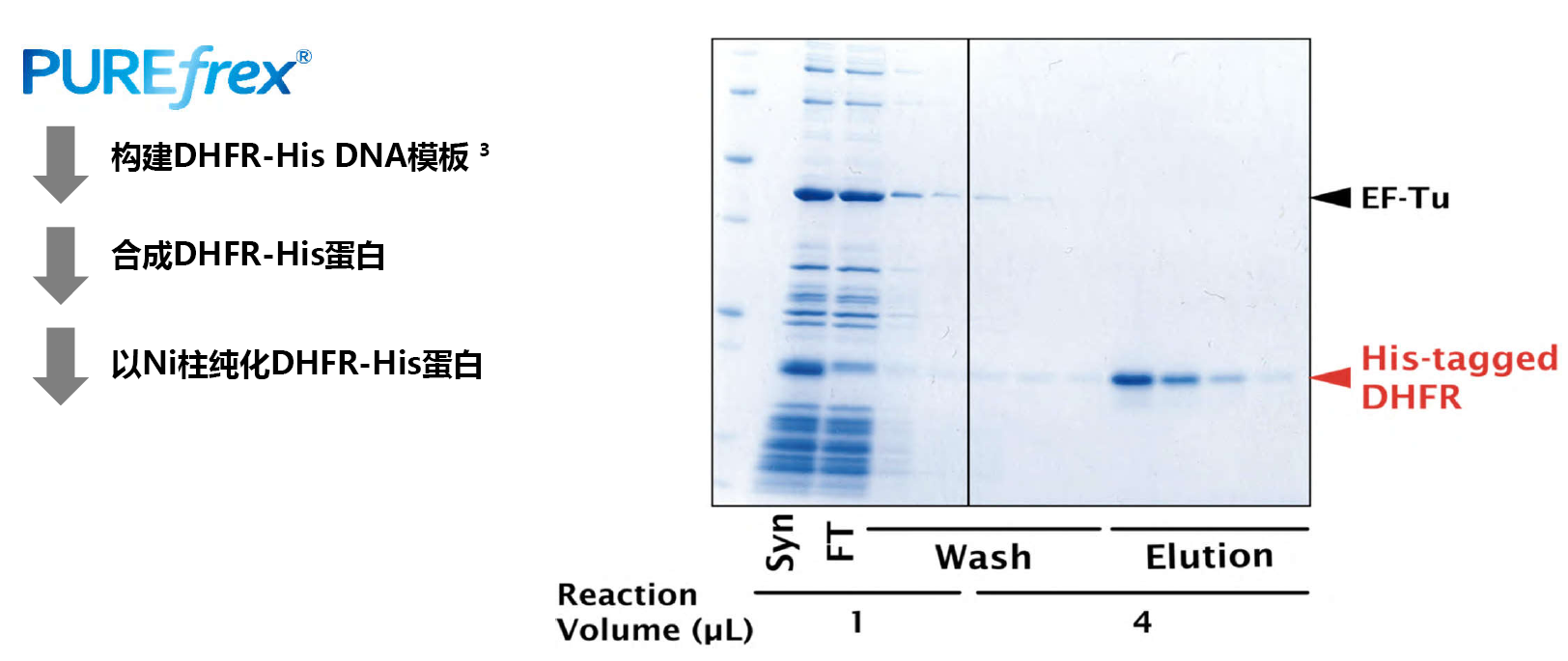
3:模板DNA的构建方法请见"相关资料"栏或点击这里。
◆添加剂(用于需要形成二硫键和分子伴侣的蛋白质)
● DS supplement
● 通过添加DS supplement至PUREfrex® 反应液中,为二硫键形成创造最佳的环境。DS supplement作为创造氧化环境的氧化剂,含有氧化型谷胱甘肽(GSSG)和作为二硫键异构酶的大肠杆菌DsbC。当蛋白需要二硫键才能产生活性形式时,请使用本添加剂。
● DnaK Mix
● DnaK Mix是高度纯化后的大肠杆菌来源的DnaK、DnaJ、GrpE蛋白以适当的浓度比例预混后的溶液。在PUREfrex® 反应体系中单独或添加DS supplement合成蛋白时同时添加DnaK Mix,可以更易获得难以独自形成高级结构的活性蛋白。
● GroE Mix
● GroE Mix是高度纯化后的大肠杆菌来源的GroEL、GroES蛋白以适当的浓度比例预混后的溶液。以PUREfrex® 反应体系合成蛋白时添加GroE Mix,可以更易获得难以独自形成高级结构的活性蛋白。
◆试剂盒组成
用于250 μL反应
使用前请将试剂盒置于-80°C保存
| 试剂 | 体积 | 成分说明 | 保存温度 |
| 溶液 I (白盖) | 125 μL | 氨基酸,核苷酸,tRNA和酶的底物等 | -20°C |
| 溶液 II (黑盖) | 12.5 μL | 蛋白,保存于含30%甘油的缓冲液 | -20°C or -80°C(1) |
| 溶液 III (红盖) | 12.5 μL ×2 | 核糖体(20 μM) | -80°C(1) |
|
DHFR DNA (透明盖)(2) |
10 μL | 对照DNA,含有编码大肠杆菌DHFR基因的PCR产物(20 ng/μL) | -20°C |
(1)剩余的溶液应快速在液氮、干冰或乙醇中冻结,并储存于-80℃。如有必要,分装剩余溶液,并尽可能避免反复冻融。
(2)每50 μL反应中加入2.5 μL DHFR DNA。
◆产品列表
|
产品编号 |
产品名称 |
规格 |
备注信息 |
|
GFK-PF201-0.25-EX |
PUREfrex® 2.0 |
1 kit |
供250 μL反应使用 |
|
GFK-PF201-0.25-5-EX |
PUREfrex® 2.0 |
1 kit |
供250 μL×5次反应使用 |
|
GFK-PF213-0.25-EX |
PUREfrex® 2.1 |
1 kit |
供250 μL反应使用 |
|
GFK-PF213-0.25-5-EX |
PUREfrex® 2.1 |
1 kit |
供250 μL×5次反应使用 |
|
GFK-PF003-0.5-EX |
DnaK Mix |
1 kit |
供500 μL反应使用 |
|
GFK-PF004-0.5-EX |
GroE Mix |
1 kit |
供500 μL反应使用 |
|
GFK-PF005-0.5-EX |
DS supplement |
1 kit |
供500 μL反应使用 |
相关产品的详细信息请点击查看:重组无细胞蛋白合成系统 PUREfrex® 2.0
相关资料
 |
 |
|
PUREfrex™ Technical information |
PUREfrex™ Protocol |
蛋白质工程相关产品
PUREfrex:重组无细胞蛋白合成试剂盒
RYTS试剂盒:大肠杆菌无细胞蛋白质合成系统
CloverDirect:定点蛋白质功能化tRNA试剂
纯化系统:一步高纯度标记纯化系统
STELLA +“赖氨酸标记试剂盒”
-
PUREfrex : Reconstituted Cell-free Protein Synthesis Kit
-
RYTS Kit : E. coli Cell-free Protein Synthesis System
-
CloverDirect : tRNA Reagents for Site-Directed Protein Functionalization
-
Dock Purification System : One step high purity purification tag purification system
-
STELLA+ " Lysine Labeling Kit "
PUREfrex™ Q&A
|
Q: 使用PUREfrex™ 试剂盒是否可用于真核蛋白的合成? A: PUREfrex™ 是由E.coli的核糖体和翻译因子组成的体外重组蛋白合成试剂盒,但也可以合成哺乳动物和植物的蛋白。目标蛋白的合成效率取决于编码蛋白的核苷酸序列,比如GC含量,稀有密码子的含量。
Q: 使用PUREfrex™ 试剂盒可以合成多少蛋白? A: 这个取决于目标蛋白。来自E.coli的二氢叶酸还原酶每毫升反应液可合成150ug。
Q: 是否可以合成大于100kDa的蛋白? A: 我们用该试剂盒合成了116kDa的蛋白。
Q: 是否可以推荐PUREfrex™ 的反应条件? A: 推荐用该试剂盒在37℃反应2~4小时。
Q: 是否可以合成和纯化标签蛋白? A: 可以使用任何标签,PUREfrex™ 试剂盒的所有蛋白成分都没有用于纯化或者检测的标签。比如,合成后可用金属螯合的树脂纯化带有His标签的目标蛋白。
Q: 合成蛋白是否经糖基化或者磷酸化修饰? A: 不。不会发生翻译后修饰,PUREfrex™ 试剂盒只是由翻译因子组成。
Q: PUREfrex™ 试剂盒是否含有分子伴侣? A: 不。PUREfrex™ 试剂盒不含有任何分子伴侣,但你可以添加分子伴侣,比如Hsp70。你可以自己制备。
Q: 用PUREfrex™ 试剂盒是否可合成含有二硫键的蛋白? A: 不行。目标蛋白合成不带有二硫键,因为翻译反应时有还原剂DTT。大多数需要二硫键才有活性的蛋白,会没有活性。
Q: PUREfrex™ 是否可合成膜蛋白? A: 一般情况,合成膜蛋白会形成聚集。为了获得能够插入到脂双层的膜蛋白,需要在合成膜蛋白时添加脂质体到PUREfrex™。
Q: 是否可合成带有[35S] 甲硫氨酸或者 [3H] 亮氨酸的蛋白? A: 添加放射性元素标记的氨基酸可以合成放射性元素标记的蛋白,比如[35S] 甲硫氨酸或者 [3H] 亮氨酸。PUREfrex™ 含有20种天然的氨基酸,浓度都在0.5mM。请优化条件。
Q: 除了T7启动子外,是否可用其他启动子? A: 我们推荐使用T7启动子的模板DNA,因为PUREfrex™ 含有转录的RNA聚合酶。当你使用其他聚合酶,制备的模板DNA要有相应聚合酶的合适启动子。
Q: 使用DHFR DNA(阳性对照)无法获得DHFR。 A: 该试剂盒由于某些原因失活。为了避免失活,请将该试剂盒存放在适当稳定。可进行分装,避免反复冻融影响试剂盒的使用效果。或者改试剂盒被核酸酶污染了。请使用不含核酸酶的水,试剂和材料。
Q: 使用试剂盒的DHFR可以得到DHFR。但是不能得到目标蛋白,或者目标蛋白量很低。 A: 1)改试剂盒由于某些原因失活了。为了避免失活,请将该试剂盒存放在适当的温度并且进行分装(避免反复冻融) A: 2)可以受核酸酶污染。为了避免核酸酶污染,请使用不含核酸酶的水,试剂和材料。 A: 3)制备的DNA模板不准确。需要制备含有T7启动子,核糖体结合位点,起始密码子,终止密码子的DNA模板。 A: 4)转录的二级结构会阻止翻译反应。这种情况,请优化模板的顺序,解决二级结构的问题。 |
| [1] | Murakami, S., Matsumoto, R., & Kanamori, T.. (2019). Constructive approach for synthesis of a functional IgG using a reconstituted cell-free protein synthesis system. Scientific reports 9(1), 671. |
| [2] | Doerr, A., de Reus, E., van Nies, P., van der Haar, M., Wei, K., Kattan, J., et al. (2019). Modelling cell-free RNA and protein synthesis with minimal systems. Physical biology, 16, 025001. |
| [3] | Dopp, J., Tamiev, D., & Reuel, N. F.. (2019). Cell-free supplement mixtures: Elucidating the history and biochemical utility of additives used to support in vitro protein synthesis in E. coli extract. Biotechnology advances, 37(1), 246-258. |
| [4] | Marsden, A. P., Hollins, J. J., O’Neill, C., Ryzhov, P., Higson, S., Mendonça, C. A., et al. (2018). Investigating the Effect of Chain Connectivity on the Folding of a Beta-Sheet Protein On and Off the Ribosome. Journal of molecular biology, 430, 5207-5216. |
| [5] | Tian, P., Steward, A., Kudva, R., Su, T., Shilling, P. J., Nickson, A. A., et al. (2018). The Folding Pathway of an Ig Domain is Conserved On and Off the Ribosome. Proceedings of the National Academy of Sciences, 201810523., 115(48), E11284-E11293. |
| [6] | Gessesse, B., Nagaike, T., Nagata, K., Shimizu, Y., & Ueda, T.. (2018). G-Protein Coupled Receptor Protein Synthesis on a Lipid Bilayer Using a Reconstituted Cell-Free Protein Synthesis System. Life, 8(4), 54. |
| [7] | Kamiya, N., Ohama, Y., Minamihata, K., Wakabayashi, R., & Goto, M.. (2018). Liquid Marbles as an Easy‐to‐Handle Compartment for Cell‐Free Synthesis and In Situ Immobilization of Recombinant Proteins. Biotechnology journal,13(12). |
| [8] | Hayase, G., & Nomura, S. I. M.. (2018). Large-Scale Preparation of Giant Vesicles by Squeezing a Lipid-Coated Marshmallow-like Silicone Gel in a Buffer. Langmuir, 34(37), 11021-11026. |
| [9] | Fujiwara, K., Ito, K., & Chiba, S.. (2018). MifM-instructed translation arrest involves nascent chain interactions with the exterior as well as the interior of the ribosome. Scientific reports, 8(1), 10311. |
| [10] | Sugimoto, S., Arita-Morioka, K. I., Terao, A., Yamanaka, K., Ogura, T., & Mizunoe, Y.. (2018). Multitasking of Hsp70 chaperone in the biogenesis of bacterial functional amyloids. Communications Biology, 1(1), 52. |
| [11] | Kamiya, Y., Arimura, Y., Ooi, H., Kato, K., Liang, X. G., & Asanuma, H.. (2018). Development of Visible‐Light‐Responsive RNA Scissors Based on a 10–23 DNAzyme. ChemBioChem. 19, 1305-1311. |
| [12] | Fujii, S., Sawa, T., Motohashi, H., & Akaike, T.. (2018). Persulfide synthases that are functionally coupled with translation mediate sulfur respiration in mammalian cells. British Journal of Pharmacology, 176(4), 607-615. |
| [13] | Komura, R., Aoki, W., Motone, K., Satomura, A., & Ueda, M.. (2018). High-throughput evaluation of T7 promoter variants using biased randomization and DNA barcoding. PLOS ONE, 13(5), e0196905. |
| [14] | van Nies, P., Westerlaken, I., Blanken, D., Salas, M., Mencía, M., & Danelon, C.. (2018). Self-replication of DNA by its encoded proteins in liposome-based synthetic cells. Nature communications, 9(1), 1583. |
| [15] | Furusato, T., Horie, F., Matsubayashi, H. T., Amikura, K., Kuruma, Y., & Ueda, T.. (2018). De novo synthesis of basal bacterial cell division proteins FtsZ, FtsA, and ZipA inside giant vesicles. ACS synthetic biology, 7(4), 953-961. |
| [16] | Natan, E., Endoh, T., Haim-Vilmovsky, L., Flock, T., Chalancon, G., Hopper, J. T., et al. (2018). Cotranslational protein assembly imposes evolutionary constraints on homomeric proteins. Nature structural & molecular biology, 25(3), 279. |
| [17] | Ito, N., Katoh, K., Kushige, H., Saito, Y., Umemoto, T., Matsuzaki, Y., et al. (2018). Ribosome incorporation into somatic cells promotes lineage transdifferentiation towards multipotency. Scientific reports, 8(1), 1634. |
| [18] | Reyes, S. G., Kuruma, Y., & Tsuda, S.. (2017). Uncovering cell-free protein expression dynamics by a promoter library with diverse strengths. bioRxiv, 214593. |
| [19] | Katano, Y., Li, T., Baba, M., Nakamura, M., Ito, M., Kojima, K., et al. (2017). Generation of thermostable Moloney murine leukemia virus reverse transcriptase variants using site saturation mutagenesis library and cell-free protein expression system. Bioscience, biotechnology, and biochemistry, 81(12), 2339-2345. |
| [20] | Chadani, Y., Niwa, T., Izumi, T., Sugata, N., Nagao, A., Suzuki, T., et al. (2017). Intrinsic ribosome destabilization underlies translation and provides an organism with a strategy of environmental sensing. Molecular cell, 68(3), 528-539. |
| [21] | Akaike, T., Ida, T., Wei, F. Y., Nishida, M., Kumagai, Y., Alam, M. M., et al. (2017). Cysteinyl-tRNA synthetase governs cysteine polysulfidation and mitochondrial bioenergetics. Nature communications, 8(1), 1177. |
| [22] | Shepherd, T. R., Du, L., Liljeruhm, J., Wang, J., Sjödin, M. O., Wetterhall, M., et al. (2017). De novo design and synthesis of a 30-cistron translation-factor module. Nucleic acids research, 45(18), 10895-10905. |
| [23] | Matsumoto, K. I., Yamazaki, K., Kawakami, S., Miyoshi, D., Ooi, T., Hashimoto, S., & Taguchi, S.. (2017). In vivo target exploration of apidaecin based on Acquired Resistance induced by Gene Overexpression (ARGO assay). Scientific reports, 7(1), 12136. |
| [24] | Judd, J., Boucher, N., Van Roey, E., Gray, T. A., & Derbyshire, K. M.. (2017). Application of distributive conjugal DNA transfer in Mycobacterium smegmatis to establish a genome-wide synthetic genetic array. Journal of Bacteriology, 199(20). |
| [25] | Goto, Y., Murakami, H., & Suga, H.. (2008). Initiating translation with D-amino acids. RNA, 14(7), 1390–1398. |
| [26] | Ueta, M., Wada, C., Bessho, Y., Maeda, M., & Wada, A.. (2017). Ribosomal protein L31 in Escherichia coli contributes to ribosome subunit association and translation, whereas short L31 cleaved by protease 7 reduces both activities. Genes to Cells, 22(5), 452-471. |
| [27] | Nilsson, O. B., Nickson, A. A., Hollins, J. J., Wickles, S., Steward, A., Beckmann, R., et al. (2017). Cotranslational folding of spectrin domains via partially structured states. Nature structural & molecular biology, 24(3), 221. |
| [28] | Fan, Y., Hoshino, T., & Nakamura, A.. (2017). Identification of a VapBC toxin–antitoxin system in a thermophilic bacterium Thermus thermophilus HB27. Extremophiles, 21(1), 153-161. |
| [29] | Scott, A., Noga, M. J., de Graaf, P., Westerlaken, I., Yildirim, E., & Danelon, C.. (2016). Cell-free phospholipid biosynthesis by gene-encoded enzymes reconstituted in liposomes. PloS one, 11(10), e0163058. |
| [30] | Nakayama, M., Komiya, S., Fujiwara, K., Horisawa, K., & Doi, N.. (2016). In vitro selection of bispecific diabody fragments using covalent bicistronic DNA display. Biochemical and biophysical research communications, 478(2), 606-611. |
| [31] | Shimizu, Y., Inoue, A., Tomari, Y., Suzuki, T., & Ueda, T.. (2001). Cell-free translation reconstituted with purified components. Nature Biotechnology, 19(8), 751-755. |
| [32] | Radomska, K. A., Ordoñez, S. R., Wösten, M. M., Wagenaar, J. A., & van Putten, J. P.. (2016). Feedback control of Campylobacter jejuni flagellin levels through reciprocal binding of FliW to flagellin and the global regulator CsrA. Molecular microbiology, 102(2), 207-220. |
| [33] | Nilsson, O. B., Müllerlucks, A., Kramer, G., Bukau, B., & Heijne, G. V.. (2016). Trigger factor reduces the force exerted on the nascent chain by a cotranslationally folding protein. Journal of Molecular Biology, 428(6), 1356-1364. |
| [34] | Chadani, Y., Niwa, T., Chiba, S., Taguchi, H., & Ito, K.. (2016). Integrated in vivo and in vitro nascent chain profiling reveals widespread translational pausing. Proceedings of the National Academy of Sciences, 113(7), E829–E838. |
| [35] | Ando, M., Akiyama, M., Okuno, D., Hirano, M., Ide, T., Sawada, S., et al. (2016). Liposome chaperon in cell-free membrane protein synthesis: one-step preparation of KcsA-integrated liposomes and electrophysiological analysis by the planar bilayer method. Biomaterials science, 4(2), 258-264. |
| [36] | Shiraishi, A., Mochizuki, S., Miyakoshi, A., Kojoh, K., & Okada, Y.. (2016). Development of human neutralizing antibody to ADAMTS4 (aggrecanase-1) and ADAMTS5 (aggrecanase-2). Biochemical and biophysical research communications, 469(1), 62-69. |
| [37] | Nagumo, Y., Fujiwara, K., Horisawa, K., Yanagawa, H., & Doi, N.. (2015). PURE mRNA display for in vitro selection of single-chain antibodies. The Journal of Biochemistry, 159(5), 519-526. |
| [38] | Niwa, T., Sasaki, Y., Uemura, E., Nakamura, S., Akiyama, M., Ando, M.,et al. (2015). Comprehensive study of liposome-assisted synthesis of membrane proteins using a reconstituted cell-free translation system. Scientific reports, 5(1), 18025. |
| [39] | Yamamoto, H., Shima, T., Yamaguchi, M., Mochizuki, Y., Hoshida, H., Kakuta, S.,et al. (2015). The thermotolerant yeast Kluyveromyces marxianus is a useful organism for structural and biochemical studies of autophagy. Journal of Biological Chemistry, 290(49), 29506–29518. |
| [40] | Ishii, E., Chiba, S., Hashimoto, N., Kojima, S., Homma, M., Ito, K., et al. (2015). Nascent chain-monitored remodeling of the Sec machinery for salinity adaptation of marine bacteria.Proceedings of the National Academy of Sciences, 112(40), E5513-E5522. |
| [41] | Nilsson, O. B., Hedman, R., Marino, J., Wickles, S., Bischoff, L., Johansson, M., et al. (2015). Cotranslational protein folding inside the ribosome exit tunnel. Cell reports, 12(10), 1533-1540. |
| [42] | Kuruma, Y., & Ueda, T.. (2016). Corrigendum: the pure system for the cell-free synthesis of membrane proteins. Nature Protocols, 11(3), 616. |
| [43] | Morita, M., Onoe, H., Yanagisawa, M., Ito, H., Ichikawa, M., Fujiwara, K., et al. (2015). Droplet‐Shooting and Size‐Filtration (DSSF) Method for Synthesis of Cell‐Sized Liposomes with Controlled Lipid Compositions. ChemBioChem, 16(14), 2029-2035. |
| [44] | Yamashita, H., Morita, M., Sugiura, H., Fujiwara, K., Onoe, H., & Takinoue, M.. (2015). Generation of monodisperse cell-sized microdroplets using a centrifuge-based axisymmetric co-flowing microfluidic device. Journal of bioscience and bioengineering, 119(4), 492-495. |
| [45] | Nies, V., & Pauline.. (2015). monitoring mrna and protein levels in bulk and in model vesicle-based artificial cells. Methods in Enzymology, 550, 187-214. |
| [46] | Ichihashi, N., Kobori, S., & Yomo, T..(2015). Simple Identification of Two Causes of Noise in an Aptazyme System by Monitoring Cell-Free Transcription. Methods in Enzymology, 550, 93-107. |
| [47] | Kogure, H., Handa, Y., Nagata, M., Kanai, N., Peter Güntert, & Kubota, K., et al. (2014). Identification of residues required for stalled-ribosome rescue in the codon-independent release factor yaej. Nucleic Acids Research, 42(5), 3152. |
| [48] | Shimizu, Y., Kuruma, Y., Kanamori, T., & Ueda, T.. (2014). The pure system for protein production. Methods in Molecular Biology, 1118(1118), 275-284. |
| [49] | Jackson, K., Kanamori, T., Ueda, T., & Fan, Z. H.. (2014). Protein synthesis yield increased 72 times in the cell-free pure system. Integrative Biology, 6(8),781-788. |
| [50] | Matsubayashi, H., Kuruma, Y., & Ueda, T.. (2014). In vitro synthesis of the e. coli sec translocon from dna. Angewandte Chemie International Edition in English, 53(29), 7535-7538. |
| [51] | Nourian, Z., Scott, A., & Danelon, C.. (2014). Toward the assembly of a minimal divisome. Systems and Synthetic Biology, 8(3), 237-247. |
| [52] | Sugimoto, N.. (2014). Noncanonical structures and their thermodynamics of dna and rna under molecular crowding: beyond the watson-crick double helix. Int Rev Cell Mol Biol, 307, 205-273. |
| [53] | Fujiwara, K., Katayama, T., & Nomura, S. I.. (2013). Cooperative working of bacterial chromosome replication proteins generated by a reconstituted protein expression system. Nucleic Acids Research, 41(14), 7176-7183. |
| [54] | Endoh, T., Kawasaki, Y., & Sugimoto, N.. (2013). Translational halt during elongation caused by g-quadruplex formed by mrna. Methods, 64(1), 73-78. |
| [55] | Hong, S. H., Ntai, I., Haimovich, A. D., Kelleher, N. L., Isaacs, F. J., & Jewett, M. C.. (2014). Cell-free protein synthesis from a release factor 1 deficient, escherichia coli, activates efficient and multiple site-specific nonstandard amino acid incorporation. ACS Synthetic Biology, 3(6), 398-409. |
| [56] | Chizzolini, F., Forlin, M., Cecchi, D., & Mansy, S. S.. (2013). Gene position more strongly influences cell-free protein expression from operons than t7 transcriptional promoter strength. ACS Synthetic Biology, 3(6). |
| [57] | Fujii, S., Matsuura, T., Sunami, T., Kazuta, Y., & Yomo, T.. (2013). In vitro evolution of -hemolysin using a liposome display. Proceedings of the National Academy of Sciences, 110(42), 16796-16801. |
| [58] | Nies, V., Pauline, Nourian, Zohreh, Kok, & Maurits, et al. (2013). Unbiased tracking of the progression of mrna and protein synthesis in; bulk and in liposome-confined reactions. Chembiochem A European Journal of Chemical Biology, 14(15), 1963-1966. |
| [59] | Niederholtmeyer, H., Stepanova, V., & Maerkl, S. J.. (2013). Implementation of cell-free biological networks at steady state. Proceedings of the National Academy of Sciences, 110(40), 15985-15990. |
| [60] | Lentini, R., Forlin, M., Martini, L., Bianco, C. D., Spencer, A. C., & Torino, D., et al. (2013). Fluorescent proteins and in vitro genetic organization for cell-free synthetic biology. ACS Synthetic Biology, 2(9), 482-489. |
| [61] | Woolstenhulme, C. J., Parajuli, S., Healey, D. W., Valverde, D. P., Petersen, E. N., & Starosta, A. L., et al. (2013). Nascent peptides that block protein synthesis in bacteria. Proceedings of the National Academy of Sciences, 110(10), E878-E887. |
| [62] | Jewett, M. C., Fritz, B. R., Timmerman, L. E., & Church, G. M.. (2014). In vitro integration of ribosomal rna synthesis, ribosome assembly, and translation. Molecular Systems Biology, 9(1), 678-678. |
| [63] | Niederholtmeyer, H., Xu, L., & Maerkl, S. J.. (2013). Real-time mrna measurementduring an in vitro transcription and translationreaction using binary probes. ACS Synthetic Biology, 2(8), 411-417. |
| [64] | Endoh, T., Kawasaki, Y., & Sugimoto, N.. (2013). Stability of rna quadruplex in open reading frame determines proteolysis of human estrogen receptor α. Nucleic Acids Research, 41(12), 6222-6231. |
| [65] | Endoh, T., Kawasaki, Y., & Sugimoto, N.. (2013). Suppression of gene expression by g-quadruplexes in open reading frames depends on g-quadruplex stability. Angewandte Chemie International Edition, 52(21), 5522-5526. |
| [66] | Lee, K. B., Kim, H. C., Kim, D. M., Kang, T. J., & Suga, H.. (2013). Comparative evaluation of two cell-free protein synthesis systems derived from escherichia coli for genetic code reprogramming. Journal of Biotechnology, 164(2), 330-335. |
| [67] | Nakamura, Y., Ogura, M., Ogura, K., Tanaka, D., & Inagaki, N.. (2012). Sirt5 deacetylates and activates urate oxidase in liver mitochondria of mice. FEBS letters, 586(23), 4076-4081. |
| [68] | Fujino, Y., Fujita, R., Wada, K., Fujishige, K., & Ueda, T.. (2012). Robust in vitro affinity maturation strategy based on interface-focused high-throughput mutational scanning. Biochemical and Biophysical Research Communications, 428(3), 395-400. |
| [69] | Venancio-Marques, A., Liu, Y.-J., Diguet, A., di Maio, T., Gautier, A., & Baigl, D. (2012). Modification-Free Photocontrol of β-Lactam Conversion with Spatiotemporal Resolution. ACS Synthetic Biology, 1(11), 526–531. |
| [70] | Nicolini, C., Bragazzi, N., & Pechkova, E.. (2012). Nanoproteomics enabling personalized nanomedicine. Advanced Drug Delivery Reviews, 64(13), 1522-1531. |
| [71] | Matsuura, T., Hosoda, K., Kazuta, Y., Ichihashi, N., Suzuki, H., & Yomo, T.. (2012). Effects of compartment size on the kinetics of intracompartmental multimeric protein synthesis. ACS Synthetic Biology, 1(9), 431-437. |
| [72] | Ong, H. J., Siau, J. W., Zhang, J. B., Hong, M., Flotow, H., & Ghadessy, F.. (2012). Analysis of p53 binding to dna by fluorescence imaging microscopy. Micron, 43(9), 996-1000. |
| [73] | Shimizu, Y.. (2012). Arfa recruits rf2 into stalled ribosomes. Journal of molecular biology, 423(4), 624-631. |
| [74] | Nagano, T., Kojima, K., Hisabori, T., Hayashi, H., Morita, E. H., & Kanamori, T., et al. (2012). Elongation factor g is a critical target during oxidative damage to the translation system of escherichia coli. Journal of Biological Chemistry, 287(34), 28697-28704. |
| [75] | Ying, & B.-W. (2003). A novel screening system for self-mrna targeting proteins. Journal of Biochemistry, 133(4), 485-491. |
| [76] | Kobori, S., Ichihashi, N., Kazuta, Y., Matsuura, T., & Yomo, T.. (2012). Kinetic analysis of aptazyme-regulated gene expression in a cell-free translation system: modeling of ligand-dependent and -independent expression. Rna-a Publication of the Rna Society, 18(8), 1458-1465. |
| [77] | Bruder, J., Siewert, K., Obermeier, B., Malotka, J., Scheinert, P., & Kellermann, J., et al. (2012). Target specificity of an autoreactive pathogenic human γδ-T cell receptor in myositis. Journal of Biological Chemistry, 287(25), 20986-20995. |
| [78] | Nishimura, K., Matsuura, T., Nishimura, K., Sunami, T., Suzuki, H., & Yomo, T.. (2012). Cell-free protein synthesis inside giant unilamellar vesicles analyzed by flow cytometry. Langmuir, 28(22), 8426-8432. |
| [79] | Okano, T., Matsuura, T., Kazuta, Y., Suzuki, H., & Yomo, T.. (2012). Cell-free protein synthesis from a single copy of dna in a glass microchamber. Lab on a Chip, 12(15), 2704. |
| [80] | Guarino, C., & Delisa, M. P.. (2012). A prokaryote-based cell-free translation system that efficiently synthesizes glycoproteins. Glycobiology, 22(5), 596-601. |
| [81] | Stögbauer, T., Windhager, L., Zimmer, R., & Rädler, J. O. (2012). Experiment and mathematical modeling of gene expression dynamics in a cell-free system. Integrative Biology, 4(5), 494-501. |
| [82] | Do, P. M., Varanasi, L., Fan, S., Li, C., Kubacka, I., & Newman, V., et al. (2012). Mutant p53 cooperates with ets2 to promote etoposide resistance. Genes & Development, 26(8), 830-845. |
| [83] | Kriechbaumer, V., Wang, P., Hawes, C., & Abell, B. M.. (2012). Alternative splicing of the auxin biosynthesis gene yucca4 determines its subcellular compartmentation. The Plant Journal, 70(2), 292-302. |
| [84] | Zhu, X., Ahmad, S. M., Aboukhalil, A., Busser, B. W., & Michelson, A. M.. (2012). Differential regulation of mesodermal gene expression by drosophila cell type-specific forkhead transcription factors. Development, 139(8), 1457-1466. |
| [85] | Guillen Schlippe, Y. V., Hartman, M. C. T., Josephson, K., & Szostak, J. W.. (2012). in vitror, selection of highly modified cyclic peptides that act as tight binding inhibitors. Journal of the American Chemical Society, 134(25), 10469-10477. |
| [86] | Takahashi, S., Tsuji, K., Ueda, T., & Okahata, Y.. (2012). Traveling time of a translating ribosome along messenger rna monitored directly on a quartz crystal microbalance. Journal of the American Chemical Society, 134(15), 6793-6800. |
| [87] | Papenfort, K., Podkaminski, D., Hinton, J. C. D., & Jörg Vogel. (2012). The ancestral sgrs rna discriminates horizontally acquired salmonella mrnas through a single g-u wobble pair. Proceedings of the National Academy of Sciences, 109(13), E757-764. |
| [88] | Danelon, C., Nourian, Z., Roelofsen, W., & Westerlaken, I.. (2012). Triggered gene expression in fed-vesicle microreactors with a multifunctional membrane. Biophysical Journal, 102(3), 715a. |
| [89] | Rosenblum, G., Chen, C., Kaur, J., Cui, X., Goldman, Y. E., & Cooperman, B. S.. (2012). Real-time assay for testing components of protein synthesis. Nucleic Acids Research, 40(12), e88-e88. |
| [90] | Machida, K., Masutani, M., Kobayashi, T., Mikami, S., Nishino, Y., & Miyazawa, A., et al. (2012). Reconstitution of the human chaperonin cct by co-expression of the eight distinct subunits in mammalian cells. Protein Expression & Purification, 82(1), 61-69. |
| [91] | Barendt, P. A., Shah, N. A., Barendt, G. A., Sarkar, C. A., & Hughes, D.. (2012). Broad-specificity mrna–rrna complementarity in efficient protein translation. PLoS Genetics, 8(3), e1002598. |
| [92] | Wang, H. H., Huang, P.-Y., Xu, G., Haas, W., Marblestone, A., Li, J. et al.. (2012). Multiplexed in Vivo His-Tagging of Enzyme Pathways for in Vitro Single-Pot Multienzyme Catalysis. ACS Synthetic Biology, 1(2), 43–52. |
| [93] | Holmqvist, E., Unoson, C., Reimegård, J., & Wagner, E. G. H. (2012). A mixed double negative feedback loop between the sRNA MicF and the global regulator Lrp. Molecular Microbiology, 84(3), 414–427. |
| [94] | Endoh, T., Kawasaki, Y., & Sugimoto, N.. (2012). Synchronized translationfor detection of temporalstalling of ribosome during single-turnover translation. Analytical Chemistry, 84(2), 857-861. |
| [95] | Marcin, D., Reynolds, C. B., & Fairweather, N. F.. (2012). Clostridium difficile cell wall protein cwpv undergoes enzyme-independent intramolecular autoproteolysis. Journal of Biological Chemistry, 287(2), 1538-1544. |
| [96] | Atsushi, O., Masayoshi, H., Shinsuke, S., & Yasuhiro, A.. (2012). A concept for selection of codon-suppressor trnas based on read-through ribosome display in an in vitro compartmentalized cell-free translation system. Journal of Nucleic Acids, 2012, 538129. |
| [97] | Lazzeriniospri, L., Stano, P., Luisi, P. L., & Marangoni, R.. (2012). Characterization of the emergent properties of a synthetic quasi-cellular system. Bmc Bioinformatics, 13(Suppl 4), S9. |
| [98] | Nobuhide, D., Natsuko, Y., Hideaki, M., Yasutsugu, Y., Tetsuya, N., & Nobutaka, M., et al. (2012). Dna display selection of peptide ligands for a full-length human g protein-coupled receptor on cho-k1 cells. PLoS ONE, 7(1), e30084. |
| [99] | Harada, R., Furumoto, S., Yoshikawa, T., Ishikawa, Y., Shibuya, K., & Okamura, N., et al. (2012). Synthesis of [11c]interleukin 8 using a cell-free translation system and l-[11c]methionine. Nuclear Medicine & Biology, 39(1), 155-160. |
| [100] | Wang, X., Morgan, R., Nugent, M. L., Gupta, Y., Xu, S., & Fomenkov, A., et al. (2011). Characterization of type ii and iii restriction-modification systems from bacillus cereus strains atcc 10987 and atcc 14579. Journal of Bacteriology, 194(1), 49-60. |
| [101] | Hufton, S. E.. (2012). Affinity maturation and functional dissection of a humanised anti-rage monoclonal antibody by ribosome display. Methods in Molecular Biology, 805, 403-422. |
| [102] | Ohashi, H., Kanamori, T., Osada, E., Akbar, B. K., & Ueda, T.. (2012). Peptide screening using pure ribosome display. Methods in Molecular Biology, 805(1), 251-259. |
| [103] | Nishikawa, T., Sunami, T., Matsuura, T., & Yomo, T. (2012). Directed Evolution of Proteins throughIn VitroProtein Synthesis in Liposomes. Journal of Nucleic Acids, 2012, 1–11. |
| [104] | Takeshi, S., Hiroshi, Y., & Nobuhide, D.. (2012). in vitro selection of fab fragments by mrna display and gene-linking emulsion pcr. Journal of Nucleic Acids, 2012, 1-9. |
| [105] | Karig, D. K., Iyer, S., Simpson, M. L., & Doktycz, M. J.. (2012). Expression optimization and synthetic gene networks in cell-free systems. Nucleic Acids Research, 40(8), 3763-3774. |
| [106] | Niwa, T., Kanamori, T., Ueda, T., & Taguchi, H..(2012). Global analysis of chaperone effects using a reconstituted cell-free translation system. Proc Natl Acad Sci USA, 109, 8937-8942. |
| [107] | Kaiser, C., Goldman, D., Tinoco, I., & Bustamante, C.. (2012). The ribosome modulates nascent protein folding. Biophysical Journal, 102(3), 68a. |
| [108] | Wang, W., Hara, S., Liu, M., Aigaki, T., Shimizu, S., & Ito, Y.. (2011). Polypeptide aptamer selection using a stabilized ribosome display. Journal of Bioscience & Bioengineering, 112(5), 515-517. |
| [109] | Gonza?Lez, D., Lokhande, N., Vadde, S., Zhao, Q., Cassill, A., & Renthal, R.. (2011). Luminescence resonance energy transfer in the cytoplasm of live escherichia coli cells. Biochemistry, 50(32), 6789-6796. |
| [110] | Mallam, A. L., & Jackson, S. E.. (2011). Knot formation in newly translated proteins is spontaneous and accelerated by chaperonins. Nature Chemical Biology, 8(2), 147-153. |
| [111] | Hensley, M. P., Tierney, D. L., & Crowder, M. W.. (2011). Zn(ii) binding to escherichia coli 70s ribosomes. Biochemistry, 50(46), 9937-9939. |
| [112] | Pereira de Souza, T., Steiniger, F., Stano, P., Fahr, A., & Luisi, P. L. (2011). Spontaneous Crowding of Ribosomes and Proteins inside Vesicles: A Possible Mechanism for the Origin of Cell Metabolism. ChemBioChem, 12(15), 2325–2330. |
| [113] | Grimm, S., Yu, F., & Nygren, P.-Å. (2011). Ribosome Display Selection of a Murine IgG1 Fab Binding Affibody Molecule Allowing Species Selective Recovery Of Monoclonal Antibodies. Molecular Biotechnology, 48(3), 263–276. |
| [114] | Yanagida, H., Matsuura, T., Kazuta, Y., & Yomo, T. (2011). In Vitro Selection of Proteins that Undergo Covalent Labeling with Small Molecules by Thiol-Disulfide Exchange by Using Ribosome Display. ChemBioChem, 12(6), 962–969. |
| [115] | Welsh, J. P., Bonomo, J., & Swartz, J. R.. (2011). Localization of bip to translating ribosomes increases soluble accumulation of secreted eukaryotic proteins in an escherichia coli cell-free system. Biotechnology & Bioengineering, 108(8), 1739-1748. |
| [116] | Kihara, F., Niimi, T., Yamashita, O., & Yaginuma, T. (2011). Heat shock factor binds to heat shock elements upstream of heat shock protein 70a and Samui genes to confer transcriptional activity in Bombyx mori diapause eggs exposed to 5°C. Insect Biochemistry and Molecular Biology, 41(11), 843–851. |
| [117] | Iizuka, R., Yamagishi-Shirasaki, M., & Funatsu, T.. (2011). Kinetic study of de novo chromophore maturation of fluorescent proteins. Biophysical Journal, 100(3), 486a. |
| [118] | Ohtsuka, T., Neki, S., Kanai, T., Akiyoshi, K., Nomura, S. M., & Ohtsuki, T.. (2011). Synthesis and in situ insertion of a site-specific fluorescently labeled membrane protein into cell-sized liposomes. Analytical Biochemistry, 418(1), 97-101. |
| [119] | Lam, K. N., Van Bakel, H., Cote, A. G., Anton, V. D. V., & Hughes, T. R.. (2011). Sequence specificity is obtained from the majority of modular c2h2 zinc-finger arrays. Nucleic Acids Research, 39(11), 4680-4690. |
| [120] | De Masi, F., Grove, C. A., Vedenko, A., Alibés, A., Gisselbrecht, S. S., Serrano, L., et al. (2011). Using a structural and logics systems approach to infer bHLH–DNA binding specificity determinants. Nucleic Acids Research, 39(11), 4553–4563. |
| [121] | Garza-Sánchez, F., Schaub, R. E., Janssen, B. D., & Hayes, C. S. (2011). tmRNA regulates synthesis of the ArfA ribosome rescue factor. Molecular Microbiology, 80(5), 1204–1219. |
| [122] | Shingaki, T., & Nimura, N.. (2011). Improvement of translation efficiency in an escherichia coli cell-free protein system using cysteine. Protein Expression & Purification, 77(2), 193-197. |
| [123] | Rosner, K., Kasprzak, M. F., Horenstein, A. C. J., Thurston, H. L., Abrams, J., & Kerwin, L. Y., et al. (2011). Engineering a waste management enzyme to overcome cancer resistance to apoptosis: adding dnase1 to the anti-cancer toolbox. Cancer Gene Therapy, 18(5), 346-357. |
| [124] | Zhou, Z. P., Shimizu, Y., Tadakuma, H., Taguchi, H., Ito, K., & Ueda, T.. (2011). Single molecule imaging of the trans-translation entry process via anchoring of the tagged ribosome. Journal of Biochemistry, 149(5), 609-618. |
| [125] | Chiba, S., Kanamori, T., Ueda, T., Akiyama, Y., Pogliano, K., & Ito, K. (2011). Recruitment of a species-specific translational arrest module to monitor different cellular processes. Proceedings of the National Academy of Sciences, 108(15), 6073–6078. |
| [126] | Yamamoto, S., Izumiya, H., Mitobe, J., Morita, M., Arakawa, E., & Ohnishi, M., et al. (2011). Identification of a chitin-induced small rna that regulates translation of the tfox gene, encoding a positive regulator of natural competence in vibrio cholerae. Journal of Bacteriology, 193(8), 1953. |
| [127] | Subtelny, A. O., Hartman, M. C. T., & Szostak, J. W. (2011). Optimal Codon Choice Can Improve the Efficiency and Fidelity of N-Methyl Amino Acid Incorporation into Peptides by In-Vitro Translation. Angewandte Chemie International Edition, 50(14), 3164–3167. |
| [128] | Handa, Yoshihiro, Inaho, Noriyuki, Nameki, & Nobukazu. (2011). Yaej is a novel ribosome-associated protein in escherichia coli that can hydrolyze peptidyl–trna on stalled ribosomes. Nucleic Acids Research, 39(5), 1739-1748. |
| [129] | Ramu, H., Nora Vázquez-Laslop, Klepacki, D., Dai, Q., & Mankin, A. S.. (2011). Nascent peptide in the ribosome exit tunnel affects functional properties of the a-site of the peptidyl transferase center. Molecular cell, 41(3), 321-330. |
| [130] | Panayiotou, C., Solaroli, N., Xu, Y., Johansson, M., & Karlsson, A.. (2011). The characterization of human adenylate kinases 7 and 8 demonstrates differences in kinetic parameters and structural organization among the family of adenylate kinase isoenzymes. Biochemical Journal, 433(3), 527. |
| [131] | Narayan, V., Pion, E., Landre, V., Muller, P., & Ball, K. L.. (2011). Docking-dependent ubiquitination of the interferon regulatory factor-1 tumor suppressor protein by the ubiquitin ligase chip. Journal of Biological Chemistry, 286(1), 607-619. |
| [132] | Lamichhane, T. N., Dinuka, A. N., Duc Anne-Cécile E., Cunningham, P. R., & Chow, C. S.. (2011). Selection of peptides targeting helix 31 of bacterial 16s ribosomal rna by screening m13 phage-display libraries. Molecules, 16(2), 1211-1239. |
| [133] | Midon, M., Schafer, P., Pingoud, A., Ghosh, M., Moon, A. F., & Cuneo, M. J., et al. (2011). Mutational and biochemical analysis of the dna-entry nuclease enda from streptococcus pneumoniae. Nucleic Acids Research, 39(2), 623-634. |
| [134] | Ma, Z., & Hartman, M. C.. (2012). In vitro selection of unnatural cyclic peptide libraries via mrna display. Methods in Molecular Biology, 805, 367-390. |
| [135] | Yamaguchi, T., Yoshinaga, N., Yazawa, T., Gen, K., & Kitano, T.. (2010). Cortisol is involved in temperature-dependent sex determination in the japanese flounder. Endocrinology, 151(8), 3900-3908. |
| [136] | Ueda, T.. (2010). Ribosome display with the pure technology. Methods in Molecular Biology, 607, 219-225. |
| [137] | Kuruma, Y., Suzuki, T., & Ueda, T.. (2010). Production of multi-subunit complexes on liposome through an e. coli cell-free expression system. Methods Mol Biol, 607, 161-171. |
| [138] | Shimizu, Y., & Ueda, T.. (2010). Pure technology. Methods in Molecular Biology, 607, 11-21. |
| [139] | Moritani, Y., Nomura, S. I. M., Morita, I., & Akiyoshi, K.. (2010). Direct integration of cell-free-synthesized connexin-43 into liposomes and hemichannel formation. Febs Journal, 277(16), 3343-3352. |
| [140] | Lakshmipathy, S. K., Gupta, R., Pinkert, S., Etchells, S. A., & Hartl, F. U.. (2010). Versatility of trigger factor interactions with ribosome-nascent chain complexes. Journal of Biological Chemistry, 285(36), 27911-27923. |
| [141] | Haruichi, A., & Shaorong, C.. (2010). In vitro genetic reconstruction of bacterial transcription initiation by coupled synthesis and detection of rna polymerase holoenzyme. Nucleic Acids Research, 38(13), e141. |
| [142] | Theerthagiri, G., Eisenhardt, N., Schwarz, H., & Antonin, W.. (2010). The nucleoporin nup188 controls passage of membrane proteins across the nuclear pore complex. The Journal of Cell Biology, 189(7), 1129-1142. |
| [143] | Shen, B. W., Heiter, D. F., Chan, S. H., Wang, H., Xu, S. Y., & Morgan, R. D., et al. (2010). Unusual target site disruption by the rare-cutting hnh restriction endonuclease paci. Structure, 18(6), 734-743. |
| [144] | Holmqvist, E., Reimeg?Rd, J., Sterk, M., Grantcharova, N., R?Mling, U., & Wagner, E. G. H.. (2010). Two antisense rnas target the transcriptional regulator csgd to inhibit curli synthesis. EMBO JOURNAL, 29(11), 1840-1850. |
| [145] | Sunami, T., Hosoda, K., Suzuki, H., Matsuura, T., & Yomo, T.. (2010). Cellular compartment model for exploring the effect of the lipidic membrane on the kinetics of encapsulated biochemical reactions. Langmuir, 26(11), 8544-8551. |
| [146] | Bonomo, J., Welsh, J. P., Manthiram, K., & Swartz, J. R.. (2010). Comparing the functional properties of the hsp70 chaperones, dnak and bip. Biophysical Chemistry, 149(1), 58-66. |
| [147] | Nishiyama, K. I., Maeda, M., Abe, M., Kanamori, T., Shimamoto, K., & Kusumoto, S., et al. (2010). A novel complete reconstitution system for membrane integration of the simplest membrane protein. Biochemical & Biophysical Research Communications, 394(3), 733-736. |
| [148] | Noto, T., Kurth, H. M., Kataoka, K., Aronica, L., Desouza, L. V., & Siu, K. W. M., et al. (2010). The tetrahymena argonaute-binding protein giw1p directs a mature argonaute-sirna complex to the nucleus. Cell, 140(5), 692-703. |
| [149] | Matsumura, N., Tsuji, T., Sumida, T., Kokubo, M., Onimaru, M., & Doi, N., et al. (2010). Mrna display selection of a high-affinity, bcl-xl-specific binding peptide. The FASEB Journal, 24(7), 2201-2210. |
| [150] | Osada, E., Shimizu, Y., Akbar, B. K., Kanamori, T., & Ueda, T.. (2009). Epitope mapping using ribosome display in a reconstituted cell-free protein synthesis system. Journal of Biochemistry, 145(5), 693-700. |
| [151] | Tanner, D. R., Cariello, D. A., Woolstenhulme, C. J., Broadbent, M. A., & Buskirk, A. R.. (2009). Genetic identification of nascent peptides that induce ribosome stalling. Journal of Biological Chemistry, 284(50), 34809-34818. |
| [152] | Sumida, T., Doi, N., & Yanagawa, H.. (2009). Bicistronic dna display for in vitro selection of fab fragments. Nucleic Acids Research, 37(22), e147. |
| [153] | Eriko, M. S., Akihiko, T., Hiroyuki, T., Takuya, M., Tsutomu, N., & Tomoji, K.. (2009). Profiling of gene-dependent translational progress in cell-free protein synthesis by real-space imaging. Analytical Biochemistry, 394(2), 275-280. |
| [154] | Yamamoto, H., Fukui, K., Takahashi, H., Kitamura, S., Shiota, T., & Terao, K., et al. (2009). Roles of tom70 in import of presequence-containing mitochondrial proteins. Journal of Biological Chemistry, 284(46), 31635-31646. |
| [155] | Göckler, N., Jofre, G., Papadopoulos, C., Soppa, U., Tejedor, F. J., & Becker, W.. (2009). Harmine specifically inhibits protein kinase DYRK1A and interferes with neurite formation. FEBS Journal, 276(21), 6324–6337. |
| [156] | Uchida, I., Ishihara, R., Tanaka, K., Hata, E., Makino, S., & Kanno, T., et al. (2009). Salmonella enterica serotype typhimurium dt104 arta-dependent modification of pertussis toxin-sensitive g proteins in the presence of [32p]nad. Microbiology, 155(11), 3710-3718. |
| [157] | Feng, Y., & Cronan, J. E.. (2009). A new member of the escherichia coli fad regulon: transcriptional regulation of fadm (ybaw). Journal of Bacteriology, 191(20), 6320-6328. |
| [158] | Solaroli, N., Panayiotou, C., Johansson, M., & Karlsson, A.. (2009). Identification of two active functional domains of human adenylate kinase 5. Febs Letters, 583(17), 2872-2876. |
| [159] | Pfeiffer, V., Papenfort, K., Lucchini, S., Hinton, J. C. D., & Vogel, J.. (2009). Coding sequence targeting by micc rna reveals bacterial mrna silencing downstream of translational initiation. NATURE STRUCTURAL & MOLECULAR BIOLOGY, 16(8), 840-846. |
| [160] | Estevez-Torres, A., Crozatier, C., Diguet, A., Hara, T., Saito, H., & Yoshikawa, K., et al. (2009). Sequence-independent and reversible photocontrol of transcription/expression systems using a photosensitive nucleic acid binder. Proceedings of the National Academy of Sciences, 106(30), 12219-12223. |
| [161] | Estevez-Torres, A., Crozatier, C., Diguet, A., Hara, T., Saito, H., & Yoshikawa, K., et al. (2009). Sequence-independent and reversible photocontrol of transcription/expression systems using a photosensitive nucleic acid binder. Proceedings of the National Academy of Sciences, 106(30), 12219-12223. |
| [162] | Takahashi, S., Iida, M., Furusawa, H., Shimizu, Y., Ueda, T., & Okahata, Y.. (2009). Real-time monitoring of cell-free translation on a quartz-crystal microbalance. Journal of the American Chemical Society, 131(26), 9326-9332. |
| [163] | Kuroha, K., Horiguchi, N., Aiba, H., & Inada, T. (2009). Analysis of nonstop mRNA translation in the absence of tmRNA inEscherichia coli. Genes to Cells, 14(6), 739–749. |
| [164] | Osada, E., Shimizu, Y., Akbar, B. K., Kanamori, T., & Ueda, T.. (2009). Epitope mapping using ribosome display in a reconstituted cell-free protein synthesis system. Journal of Biochemistry, 145(5), 693-700. |
| [165] | Niwa, T., Ying, B. W., Saito, K., Jin, W., Takada, S., & Ueda, T., et al. (2009). Bimodal protein solubility distribution revealed by an aggregation analysis of the entire ensemble of escherichia coli proteins. Proceedings of the National Academy of Sciences, 106(11), 4201-4206. |
| [166] | Robin, Togashi, S., Ryder, D. M., Wall, A. G., & J., G.. (2009). Trigger factor from the psychrophilic bacterium psychrobacter frigidicola is a monomeric chaperone. Journal of Bacteriology, 191(4), 1162-1168. |
| [167] | Matsuura, T., Kazuta, Y., Aita, T., Adachi, J., & Yomo, T.. (2009). Quantifying epistatic interactions among the components constituting the protein translation system. Molecular Systems Biology, 5(1). |
| [168] | Zheng, Y., Posfai, J., Morgan, R. D., Vincze, T., & Roberts, R. J.. (2009). Using shotgun sequence data to find active restriction enzyme genes. Nucleic Acids Research, 37(1), e1. |
| [169] | Hosoda, K., Sunami, T., Kazuta, Y., Matsuura, T., Suzuki, H., & Yomo, T.. (2008). Quantitative study of the structure of multilamellar giant liposomes as a container of protein synthesis reaction. Langmuir, 24(23), 13540-13548. |
| [170] | Terashima, H., Abe-Yoshizumi, R., Kojima, S., & Homma, M.. (2008). Cell-free synthesis of the torque-generating membrane proteins, poma and pomb, of the na+-driven flagellar motor in vibrio alginolyticus. Journal of Biochemistry, 144(5), 635-642. |
| [171] | Kazuta, Y., Adachi, J., Matsuura, T., Ono, N., Mori, H., & Yomo, T.. (2008). Comprehensive analysis of the effects of escherichia coli orfs on protein translation reaction. Molecular & Cellular Proteomics, 7(8), 1530-1540. |
| [172] | Maki, K., Uno, K., Morita, T., & Aiba, H.. (2008). Rna, but not protein partners, is directly responsible for translational silencing by a bacterial hfq-binding small rna. Proceedings of the National Academy of Sciences, 105(30), 10332-10337. |
| [173] | Uemura, S., Iizuka, R., Ueno, T., Shimizu, Y., Taguchi, H., & Ueda, T., et al. (2008). Single-molecule imaging of full protein synthesis by immobilized ribosomes. Nucleic Acids Research, 36(12), e70. |
| [174] | Uemura, S., Iizuka, R., Ueno, T., Shimizu, Y., Taguchi, H., & Ueda, T., et al. (2008). Single-molecule imaging of full protein synthesis by immobilized ribosomes. Nucleic Acids Research, 36(12), e70. |
| [175] | Sako, Y., Morimoto, J., Murakami, H., & Suga, H.. (2008). Ribosomal synthesis of bicyclic peptides via two orthogonal inter-side-chain reactions. Journal of the American Chemical Society, 130(23), 7232-7234. |
| [176] | Vazquezlaslop, N., Thum, C., & Mankin, A. S.. (2008). Molecular mechanism of drug-dependent ribosome stalling. Molecular Cell, 30(2), 190-202. |
| [177] | Sako, Y., Goto, Y., Murakami, H., & Suga, H.. (2008). Ribosomal synthesis of peptidase-resistant peptides closed by a nonreducible inter-side-chain bond. ACS Chemical Biology, 3(4), 241-249. |
| [178] | Urban, J. H., & Vogel, J.. (2008). Two seemingly homologous noncoding rnas act hierarchically to activate glms mrna translation. PLoS Biology, 6(3), e64. |
| [179] | Ozaki, Y., Suzuki, T., Kuruma, Y., Ueda, T., & Yoshida, M.. (2008). Unci protein can mediate ring-assembly of c-subunits of fof1-atp synthase in vitro. Biochemical & Biophysical Research Communications, 367(3), 663-666. |
| [180] | Sakamoto, A., Yamagishi, M., Watanabe, T., Aizawa, Y., Kato, T., & Funatsu, T.. (2008). Fluorescence labeling of a cytokine with desthiobiotin-tagged fluorescent puromycin. Journal of Bioscience & Bioengineering, 105(3), 238-242. |
| [181] | Goto, Y., Ohta, A., Sako, Y., Yamagishi, Y., Murakami, H., & Suga, H.. (2008). Reprogramming the translation initiation for the synthesis of physiologically stable cyclic peptides. ACS Chemical Biology, 3(2), 120-129. |
| [182] | Kawakami, T., Murakami, H., & Suga, H.. (2008). Messenger rna-programmed incorporation of multiple n-methyl-amino acids into linear and cyclic peptides. Chemistry & Biology, 15(1), 32-42. |
| [183] | Neely, R. K., & Roberts, R. J.. (2008). The BsaHI restriction-modification system: Cloning, sequencing and analysis of conserved motifs. BMC Molecular Biology, 9(1), 48. |
| [184] | Hillebrecht, J. R., & Chong, S.. (2008). A comparative study of protein synthesis in in vitro systems: from the prokaryotic reconstituted to the eukaryotic extract-based. BMC Biotechnology, 8(1), 58. |
| [185] | Yanagida, H., Matsuura, T., & Yomo, T.. (2008). Compensatory evolution of a ww domain variant lacking the strictly conserved trp residue. Journal of Molecular Evolution, 66(1), 61-71. |
| [186] | Ohta, A., Murakami, H., Higashimura, E., & Suga, H.. (2007). Synthesis of polyester by means of genetic code reprogramming. Chemistry & Biology (Cambridge), 14(12), 1315-1322. |
| [187] | Doi, Y., Ohtsuki, T., Shimizu, Y., Ueda, T., & Sisido, M.. (2007). Elongation factor tu mutants expand amino acid tolerance of protein biosynthesis system. Journal of the American Chemical Society, 129(46), 14458-14462. |
| [188] | Murtas, G., Kuruma, Y., Bianchini, P., Diaspro, A., & Luisi, P. L. (2007). Protein synthesis in liposomes with a minimal set of enzymes. Biochemical and Biophysical Research Communications, 363(1), 12–17. |
| [189] | Sharma, C. M., Darfeuille, F., Plantinga, T. H., & Vogel, J.. (2007). A small rna regulates multiple abc transporter mrnas by targeting c/a-rich elements inside and upstream of ribosome-binding sites. Genes & Development, 21(21), 2804-2817. |
| [190] | Kojima, K., Oshita, M., Nanjo, Y., Kasai, K., Tozawa, Y., Hayashi, H., & Nishiyama, Y. (2007). Oxidation of elongation factor G inhibits the synthesis of the D1 protein of photosystem II. Molecular Microbiology, 65(4), 936–947. |
| [191] | Sando, S., Abe, K., Sato, N., Shibata, T., Mizusawa, K., & Aoyama, Y. (2007). Unexpected Preference of theE. coliTranslation System for the Ester Bond during Incorporation of Backbone-Elongated Substrates. Journal of the American Chemical Society, 129(19), 6180–6186. |
| [192] | Lakshmipathy, S. K., Tomic, S., Kaiser, C. M., Chang, H. C., Genevaux, P., & Georgopoulos, C., et al. (2007). Identification of nascent chain interaction sites on trigger factor. Journal of Biological Chemistry, 282(16), 12186-12193. |
| [193] | Matsuura, T., Yanagida, H., Ushioda, J., Urabe, I., & Yomo, T. (2007). Nascent chain, mRNA, and ribosome complexes generated by a pure translation system. Biochemical and Biophysical Research Communications, 352(2), 372–377. |
| [194] | Ohashi, H., Shimizu, Y., Ying, B. W., & Ueda, T.. (2007). Efficient protein selection based on ribosome display system with purified components. Biochemical & Biophysical Research Communications, 352(1), 270-276. |
| [195] | Udagawa, T., Shimizu, Y., & Ueda, T.. (2004). Evidence for the translation initiation of leaderless mrnas by the intact 70 s ribosome without its dissociation into subunits in eubacteria. Journal of Biological Chemistry, 279(10), 8539-8546. |
| [196] | Ueno, S., Arai, H., Suzuki, M., & Husimi, Y.. (2007). An mrna-protein fusion at n-terminus for evolutionary protein engineering. International Journal of Biological Sciences, 3(6), 365-374. |
| [197] | Narita, A., Ogawa, K., Sando, S., & Aoyama, Y.. (2007). cis-regulatory hairpin-shaped mrna encoding a reporter protein: catalytic sensing of nucleic acid sequence at single nucleotide resolution. NATURE PROTOCOLS, 2(5), 1105-1116. |
| [198] | Yoshimori, A., Sakai, J., Sunaga, S., Kobayashi, T., & Tanuma, S. I.. (2007). Structural and functional definition of the specificity of a novel caspase-3 inhibitor, ac-dnld-cho. BMC Pharmacology, 7(1), 8. |
| [199] | Kawahashi, Y., Doi, N., Oishi, Y., Tsuda, C., Takashima, H., & Baba, T., et al. (2006). High-throughput fluorescence labelling of full-length cdna products based on a reconstituted translation system. Journal of Biochemistry, 141(1), 19-24. |
| [200] | Saguy, M., Gillet, R., Skorski, P., Hermann-Le Denmat, S., & Felden, B.. (2007). Ribosomal protein s1 influences trans-translation in vitro and in vivo. Nucleic Acids Research, 35(7), 2368-2376. |
| [201] | Zheng, Y., & Roberts, R. J.. (2007). Selection of restriction endonucleases using artificial cells. Nucleic Acids Research, 35(11), e83. |
| [202] | Setoguchi, K., Otera, H., & Mihara, K.. (2006). Cytosolic factor- and tom-independent import of c-tail-anchored mitochondrial outer membrane proteins. EMBO JOURNAL, 25(24), 5635-5647. |
| [203] | Sunami, T., Sato, K., Matsuura, T., Tsukada, K., Urabe, I., & Yomo, T.. (2006). Femtoliter compartment in liposomes for in vitro selection of proteins. Analytical Biochemistry, 357(1), 128-136. |
| [204] | Ying, & B.-W. (2006). Co-translational binding of groel to nascent polypeptides is followed by post-translational encapsulation by groes to mediate protein folding. Journal of Biological Chemistry, 281(31), 21813-21819. |
| [205] | Ishihara, N., Fujita, Y., Oka, T., & Mihara, K.. (2006). Regulation of mitochondrial morphology through proteolytic cleavage of opa1. EMBO JOURNAL, 25(13), 2966-2977. |
| [206] | Groves, M., Lane, S.,Douthwaite, J., Lowne, D., Rees, D. G., & Edwards, B., et al. (2012). Affinity maturation of phage display antibody populations using ribosome display. Methods in Molecular Biology, 313(1), 129-139. |
| [207] | Villemagne, D., Jackson, R., & Douthwaite, J. A.. (2006). Highly efficient ribosome display selection by use of purified components for in vitro translation. Journal of Immunological Methods, 313(1-2), 140-148. |
| [208] | Yamamoto, T., Izumi, S., & Gekko, K.. (2006). Mass spectrometry of hydrogen/deuterium exchange in 70s ribosomal proteins from e. coli. Febs Letters, 580(15), 0-3642. |
| [209] | Shimizu, Y., & Ueda, T.. (2006). Smpb triggers gtp hydrolysis of elongation factor tu on ribosomes by compensating for the lack of codon-anticodon interaction during trans-translation initiation. Journal of Biological Chemistry, 281(23), 15987-15996. |
| [210] | Seebeck, F. P., & Szostak, J. W.. (2006). Ribosomal synthesis of dehydroalanine-containing peptides. Journal of the American Chemical Society, 128(22), 7150-7151. |
| [211] | Kubota, S., Kubota, H., & Nagata, K.. (2006). Cytosolic chaperonin protects folding intermediates of gβ from aggregation by recognizing hydrophobic β-strands. Proceedings of the National Academy of Sciences of the United States of America, 103(22), 8360-8365. |
| [212] | Muto, H., Nakatogawa, H., & Ito, K.. (2006). Genetically encoded but nonpolypeptide prolyl-trna functions in the a site for secm-mediated ribosomal stall. Molecular Cell, 22(4), 545-552. |
| [213] | Murakami, H., Ohta, A., Ashigai, H., & Suga, H.. (2006). A highly flexible trna acylation method for non-natural polypeptide synthesis. Nature Methods, 3(5), 357-359. |
| [214] | Umekage, S., & Ueda, T.. (2006). Spermidine inhibits transient and stable ribosome subunit dissociation. Febs Letters, 580(5), 0-1226. |
| [215] | Itoh, H., Kawazoe, Y., & Shiba, T.. (2006). Enhancement of protein synthesis by an inorganic polyphosphate in an e. coli cell-free system. Journal of Microbiological Methods, 64(2), 241-249. |
| [216] | Ogawa, A., Sando, S., & Aoyama, Y.. (2010). Termination‐free prokaryotic protein translation by using anticodon‐adjusted e. coli trnaser as unified suppressors of the uaa/uga/uag stop codons. read‐through ribosome display of full‐length dhfr with translated utr as a buried spacer arm. Chembiochem, 7(2), 249-252. |
| [217] | Tomic, S., Johnson, A. E., Hartl, F. U., & Etchells, S. A. (2005). Exploring the capacity of trigger factor to function as a shield for ribosome bound polypeptide chains. FEBS Letters, 580(1), 72–76. |
| [218] | Hallier, M. (2006). Small protein B interacts with the large and the small subunits of a stalled ribosome during trans-translation. Nucleic Acids Research, 34(6), 1935–1943. |
| [219] | Jarutat, T., Frisch, C., Nickels, C., Merz, H., & Knappik, A.. (2006). Isolation and comparative characterization of ki-67 equivalent antibodies from the hucal? phage display library. Biological Chemistry, 387(7). |
| [220] | Josephson, K., Hartman, M. C. T., & Szostak, J. W. (2005). Ribosomal Synthesis of Unnatural Peptides. Journal of the American Chemical Society, 127(33), 11727–11735. |
| [221] | Shimizu, Y., Kanamori, T., & Ueda, T.. (2005). Protein synthesis by pure translation systems. Methods (Amsterdam), 36(3), 299-304. |
| [222] | Sando, S., Kanatani, K., Sato, N., Matsumoto, H., Hohsaka, T., & Aoyama, Y.. (2005). A small-molecule-based approach to sense codon-templated natural-unnatural hybrid peptides. selective silencing and reassignment of the sense codon by orthogonal reacylation stalling at the single-codon level. Journal of the American Chemical Society, 127(22), 7998-7999. |
| [223] | Fukushima, K., Ikehara, Y., & Yamashita, K. (2005). Functional Role Played by the Glycosylphosphatidylinositol Anchor Glycan of CD48 in Interleukin-18-induced Interferon-γ Production. Journal of Biological Chemistry, 280(18), 18056–18062. |
| [224] | Yano, M., Okano, H. J., & Okano, H.. (2005). Involvement of hu and heterogeneous nuclear ribonucleoprotein k in neuronal differentiation through p21 mrna post-transcriptional regulation. Journal of Biological Chemistry, 280(13), 12690-12699. |
| [225] | Ying, B. W., Taguchi, H., Kondo, M., & Ueda, T.. (2005). Co-translational involvement of the chaperonin groel in the folding of newly translated polypeptides. Journal of Biological Chemistry, 280(12), 12035-12040. |
| [226] | Tokunaga, M., Mizukami, M., & Tanaka, R.. (2005). Novel processing and localization of cata, ccda associated thiol-disulfide oxidoreductase, in protein hyper-producing bacterium brevibacillus choshinensis. Protein & Peptide Letters, 12(1), 95-98. |
| [227] | Kuruma, Y., Nishiyama, K. I., Shimizu, Y., Matthias Müller, & Ueda, T.. (2005). Development of a minimal cell-free translation system for the synthesis of presecretory and integral membrane proteins. Biotechnology progress, 21(4), 1243-1251. |
| [228] | Ying, B.-W., Taguchi, H., Ueda, H., & Ueda, T. (2004). Chaperone-assisted folding of a single-chain antibody in a reconstituted translation system. Biochemical and Biophysical Research Communications, 320(4), 1359–1364. |
| [229] | Asai, T., Takahashi, T., Esaki, M., Nishikawa, S. I., Ohtsuka, K., & Nakai, M., et al. (2004). Reinvestigation of the requirement of cytosolic atp for mitochondrial protein import. Journal of Biological Chemistry, 279(19), 19464-19470. |
| [230] | Kawano, M., Suzuki, S., Suzuki, M., Oki, J., & Imamura, T.. (2004). Bulge- and basal layer-specific expression of fibroblast growth factor-13 (fhf-2) in mouse skin. Journal of Investigative Dermatology, 122(5), 1084-1090. |
Pall颇尔滤膜 4661 SYRINGE FILTER PES 32MM PK1000
Pall颇尔滤膜 4661 SYRINGE FILTER PES 32MM PK1000 1436 11488
Whatman 滤膜 80590501 Domestic PE Custom Capsules
Whatman 滤膜 80590501 Domestic PE Custom Capsules Domestic PE Custom Capsules
Whatman 滤膜 6771-1304 Puradisc 13针头式滤器, 0.45m 醋酸纤维素 100/盒
Whatman 滤膜 6771-1304 Puradisc 13针头式滤器, 0.45m 醋酸纤维素 100/盒 PURADISC 13/0.45 CA 100/PK
水溶性荧光染料43390 Sulfo-Cyanine5 carboxylic acid 25 mg
水溶性lumiprobe荧光染料43390 Sulfo-Cyanine5 carboxylic acid 25 mg 水溶性lumiprobe荧光染料Sulfo-Cyanine5 羧酸 25 mg 690
葡萄糖苷酸酶检测底物 Aldouronic Acid Mixture (Reduced) 货号:O-AMXR Megazyme中文站
葡萄糖苷酸酶检测底物
英文名:Aldouronic Acid Mixture (Reduced)
货号:O-AMXR
规格:100 mg
High purity Aldouronic Acids Mixture (NaBH4 Reduced) for use in research, biochemical enzyme assays and in vitro diagnostic analysis.
Contains a mixture of borohydride reduced aldobiouronic, aldotriouronic and aldotetraouronic and aldopentaouronic acids (10:60:20:10). Substrate for α-glucuronidase.
暂无问题解答
暂无视频
康宁corning 3990 PLT,96WL,WHT,NBS,NS,BK,5/25 96孔板 白色 平底 NBS表面 未灭菌 散装 5个/包
康宁corning 3990 PLT,96WL,WHT,NBS,NS,BK,5/25
96孔板 白色 平底 NBS表面 未灭菌 散装 5个/包 5包/箱 1429.51
Pall颇尔滤膜 4631 VACUCAP 60 0.1UM STERILE PK10
Pall颇尔滤膜 4631 VACUCAP 60 0.1UM STERILE PK10 294 2350
Pall颇尔滤膜 60170 FILTER DISC .45UM 13MM PK100
Pall颇尔滤膜 60170 FILTER DISC .45UM 13MM PK100 50 402
抗结蛋白抗体[ y66 ] – Cytoskeleton Marker(ab32362)
种属反应性
靶标
功能 结蛋白III类中间纤维在肌肉细胞中发现的。 成人横纹肌形成纤维网络连接的肌原纤维彼此的质膜从Z线结构的外围。 疾病相关 缺陷在DES是肌病肌原纤维结蛋白相关的原因(mfm-des)[ 601419 ];MIM:也称为结蛋白相关肌病(DRM)。 神经肌肉疾病,骨骼肌肉无力,心脏传导阻滞、心律失常有关,限制性心脏衰竭,和肌原纤维破坏反应结蛋白沉积在心脏和骨骼肌细胞胞浆内的积累。 缺陷在DES是心肌病引起扩张型1I(cmd1i)[ 604765 ] MIM。 扩张型心肌病是一种以心室扩张和收缩功能受损的疾病,导致充血性心力衰竭和心律失常。 患者的过早死亡风险。 缺陷在DES的神经源性scapuloperoneal综合征原因(Kaeser凯撒型综合征):[ 181400 ] MIM。 凯撒综合征是一种常染色体显性遗传性疾病,有特殊的无力和萎缩scapuloperoneal分布。 大量临床变化观察从scapuloperoneal,变的心脏或呼吸系统受累肢体远端弓鳍鱼和表型。 面瘫,常见的附加症状是吞咽困难和男性乳房发育症。 受影响的人似乎承担较高风险突发心脏死亡相比,受影响的妇女。 肌肉活检标本的组织学和免疫组织化学检查显示广泛的研究范围从接近正常或非特异性病理改变典型,肌原纤维与积累的结蛋白的变化。 序列相似性 属于中间丝家族。 细胞定位 细胞质. -
康宁corning 8396 BOTTLE,1000ML,.45MM,PS,W/CAP,S 储液瓶 传统风格 1000ml 45mm颈尺寸 带盖 灭菌 1个/包
康宁corning 8396 BOTTLE,1000ML,.45MM,PS,W/CAP,S
储液瓶 传统风格 1000ml 45mm颈尺寸 带盖 灭菌 1个/包 12包/箱 893.75
Whatman 滤膜 FPP-148L GF/BFIR 5 1/8x15IN 50/PK
Whatman 滤膜 FPP-148L GF/BFIR 5 1/8x15IN 50/PK GF/BFIR 5 1/8x15IN 50/PK


